.. raw:: html

.. image:: https://travis-ci.org/Edinburgh-Genome-Foundry/genedom.svg?branch=master
:target: https://travis-ci.org/Edinburgh-Genome-Foundry/genedom
:alt: Travis CI build status
.. image:: https://coveralls.io/repos/github/Edinburgh-Genome-Foundry/genedom/badge.svg?branch=master
:target: https://coveralls.io/github/Edinburgh-Genome-Foundry/genedom?branch=master
Genedom is a python library for managing the domestication of genetic parts
(i.e. the modification of their sequence so as to make them compatible with a
given genetic assembly standard). Genedom binds together a
`sequence optimizer `_,
genetic standards informations, and a reporting routines, to automate the
domestication of large batches in an easy and human-friendly way.
.. raw:: html
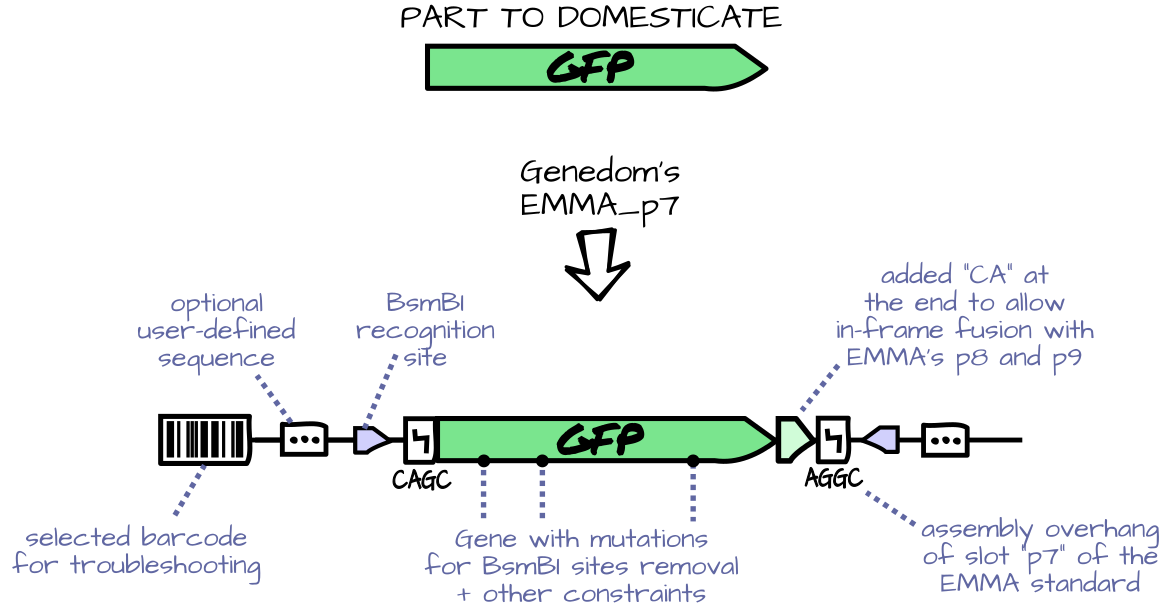
Features include:
- Possibility to define parts domesticators with added right-hand and left-hand
nucleotide, hard constraints on the sequence (such as absence of a restriction
site) and optimization objectives (such as codon optimization).
- Built-in pre-defined domesticators for popular genetic assembly standards
(well, only EMMA at the moment).
- Possibility to generate and attribute barcodes that will be added to the
sequence (but won't be in final constructs) in order to easily check
that this is the right part in the future in case of label mix-up.
- Routine for mass-domesticating sequences with report generation, including
reports on each sequence optimization, spreadsheets of parts, ready-to-order FASTA
records of the parts, and a summary report to quickly verify everything, with a
list of every domesticator used, for traceability.
Here is an example of summary report:
.. raw:: html
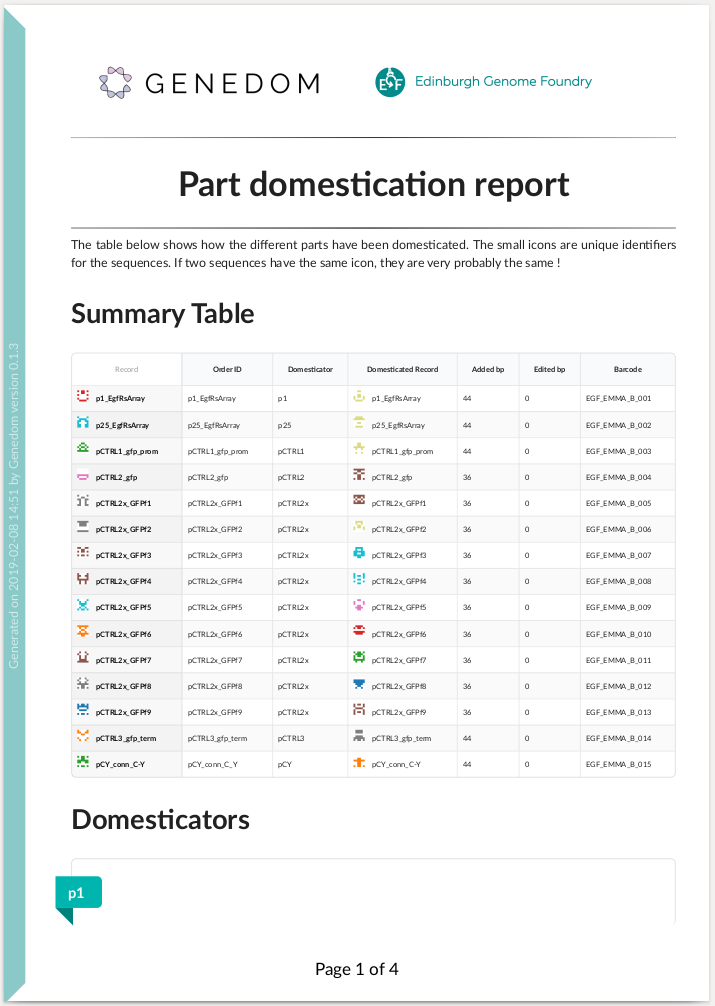
You can also use Genedom online via `this web app `_.
Usage examples
--------------
Simple domestication of one part
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
.. code:: python
from genedom import (GoldenGateDomesticator, random_dna_sequence,
write_record)
sequence = random_dna_sequence(2000, seed=123)
domesticator = GoldenGateDomesticator("ATTC", "ATCG", enzyme='BsmBI')
domestication_results = domesticator.domesticate(sequence, edit=True)
print (domestication_results.summary())
write_record(domestication_results.record_after, 'domesticated.gb')
Generating a collection of 20bp barcodes
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
(see docs for more potions)
.. code:: python
from genedom import BarcodesCollection
barcodes_collection = BarcodesCollection.from_specs(
n_barcodes=96, barcode_length=20,
forbidden_enzymes=('BsaI', 'BsmBI', 'BbsI'))
barcodes_collection.to_fasta('example_barcodes_collection.fa')
Domesticating a batch of parts with PDF report
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
.. code :: python
from genedom import BUILTIN_STANDARDS, load_record, batch_domestication
records = [
load_record(filepath, name=filename)
for filepath in records_filepaths
]
barcodes_collection = BarcodesCollection.from_specs(n_barcodes=10)
batch_domestication(records, 'domestication_report.zip',
barcodes=barcodes, # optional
standard=BUILTIN_STANDARDS.EMMA)
Installation
------------
You can install Genedom through PIP (coming soon)
.. code:: shell
sudo pip install geneblocks
Licence
-------
Genedom is an open-source software originally written at the `Edinburgh Genome Foundry
`_ by `Zulko `_
and `released on Github `_ under the MIT licence (Copyright 2018 Edinburgh Genome Foundry, University of Edinburgh).
Everyone is welcome to contribute!
More biology software
---------------------
.. image:: https://raw.githubusercontent.com/Edinburgh-Genome-Foundry/Edinburgh-Genome-Foundry.github.io/master/static/imgs/logos/egf-codon-horizontal.png
:target: https://edinburgh-genome-foundry.github.io/
Genedom is part of the `EGF Codons `_ synthetic biology software suite for DNA design, manufacturing and validation.
.. .. toctree::
.. :hidden:
.. :maxdepth: 3
.. self
.. toctree::
:hidden:
:caption: Reference
:maxdepth: 3
ref
.. _Github: https://github.com/EdinburghGenomeFoundry/genedom
.. _PYPI: https://pypi.python.org/pypi/dnachisel



