
DNA Chisel - a versatile sequence optimizer¶
DNA Chisel (complete documentation here) is a Python library for optimizing DNA sequences with respect to a set of constraints and optimization objectives. It can also be used via a command-line interface, or a web application.
The library comes with over 15 classes of sequence specifications which can be composed to, for instance, codon-optimize genes, meet the constraints of a commercial DNA provider, avoid homologies between sequences, tune GC content, or all of this at once! Users can also define their own specifications using Python, making the library suitable for a large range of automated sequence design applications, and complex custom design projects. A specification can be either a hard constraint, which must be satisfied in the final sequence, or an optimization objective, whose score must be maximized. For more information, please see the publication.
Citation¶
DNA Chisel, a versatile sequence optimizer, Valentin Zulkower, Susan Rosser. Bioinformatics (2020) 36, 16, 4508–4509
Usage¶
Defining a problem via scripts¶
The example below will generate a random sequence and optimize it so that:
It will be rid of BsaI sites (on both strands).
GC content will be between 30% and 70% on every 50bp window.
The reading frame at position 500-1400 will be codon-optimized for E. coli.
from dnachisel import *
# DEFINE THE OPTIMIZATION PROBLEM
problem = DnaOptimizationProblem(
sequence=random_dna_sequence(10000),
constraints=[
AvoidPattern("BsaI_site"),
EnforceGCContent(mini=0.3, maxi=0.7, window=50),
EnforceTranslation(location=(500, 1400))
],
objectives=[CodonOptimize(species='e_coli', location=(500, 1400))]
) # Note: always use a codon optimisation specification with EnforceTranslation
# SOLVE THE CONSTRAINTS, OPTIMIZE WITH RESPECT TO THE OBJECTIVE
problem.resolve_constraints()
problem.optimize()
# PRINT SUMMARIES TO CHECK THAT CONSTRAINTS PASS
print(problem.constraints_text_summary())
print(problem.objectives_text_summary())
# GET THE FINAL SEQUENCE (AS STRING OR ANNOTATED BIOPYTHON RECORDS)
final_sequence = problem.sequence # string
final_record = problem.to_record(with_sequence_edits=True)
Defining a problem via Genbank features¶
You can also define a problem by annotating directly a Genbank as follows:
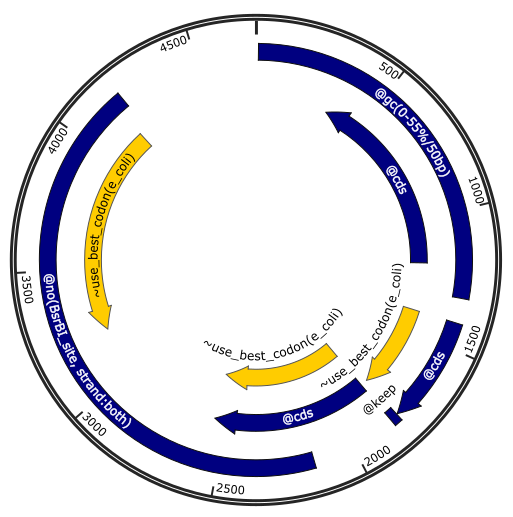
Note that constraints (colored in blue in the illustration) are features of type
misc_feature with a prefix @ followed by the name of the constraints
and its parameters, which are the same as in python scripts. Optimization
objectives (colored in yellow in the illustration) use prefix ~. See
the Genbank API documentation
for more details.
Genbank files with specification annotations can be directly fed to the web application or processed via the command line interface:
# Output the result to "optimized_record.gb"
dnachisel annotated_record.gb optimized_record.gb
Or via a Python script:
from dnachisel import DnaOptimizationProblem
problem = DnaOptimizationProblem.from_record("my_record.gb")
problem.optimize_with_report(target="report.zip")
By default, only the built-in specifications of DNA Chisel can be used in the annotations, however it is easy to add your own specifications to the Genbank parser, and build applications supporting custom specifications on top of DNA Chisel.
Reports¶
DNA Chisel also implements features for verification and troubleshooting. For instance by generating optimization reports:
problem = DnaOptimizationProblem(...)
problem.optimize_with_report(target="report.zip")
Here is an example of summary report:
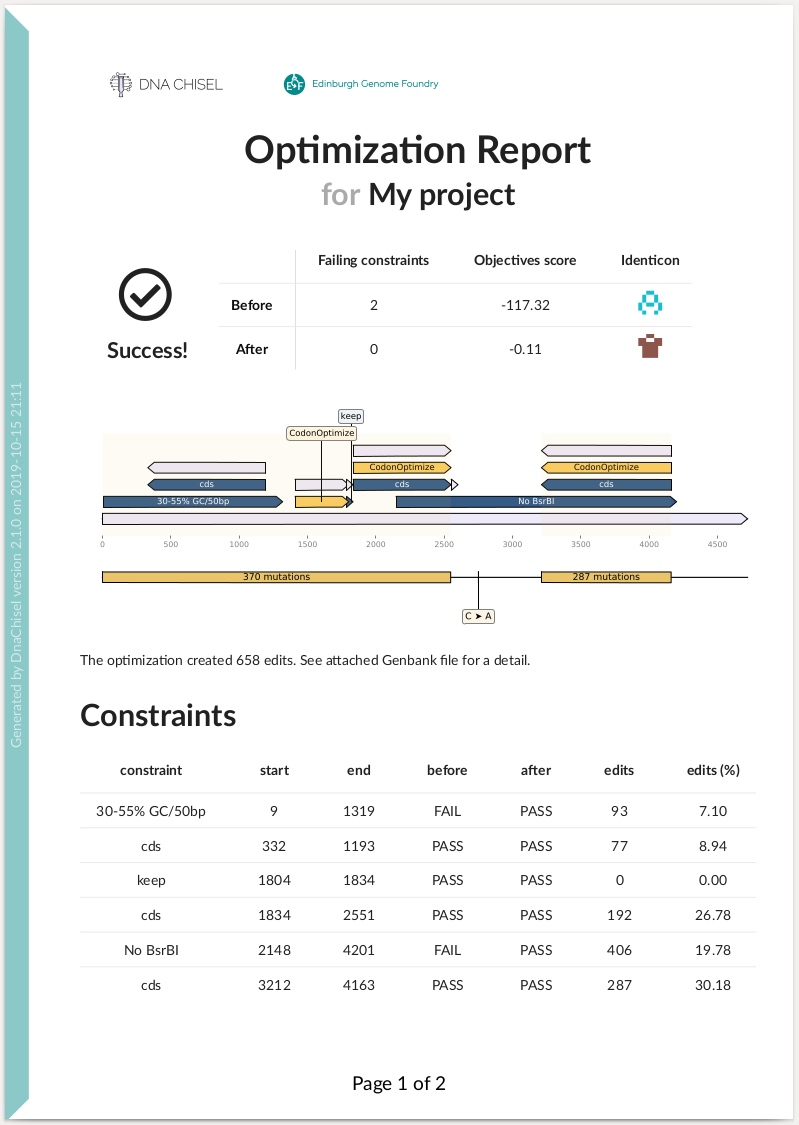
How it works¶
DNA Chisel hunts down every constraint breach and suboptimal region by recreating local version of the problem around these regions. Each type of constraint can be locally reduced and solved in its own way, to ensure fast and reliable resolution.
Below is an animation of the algorithm in action:
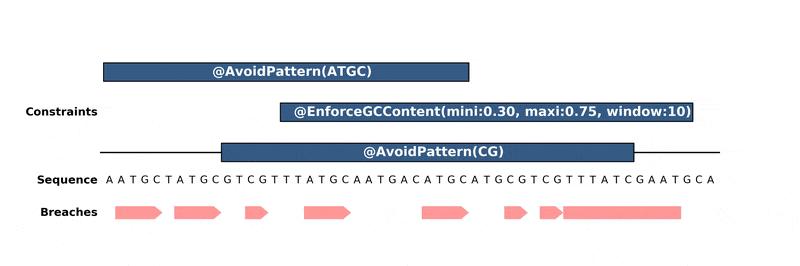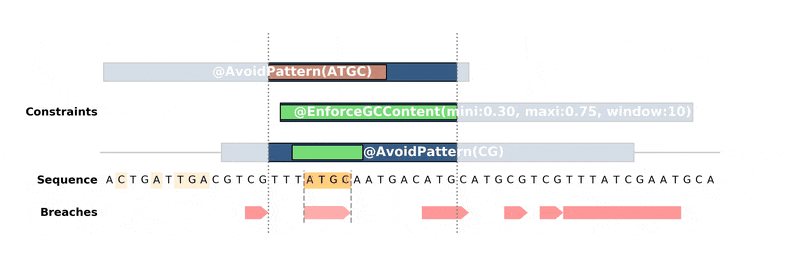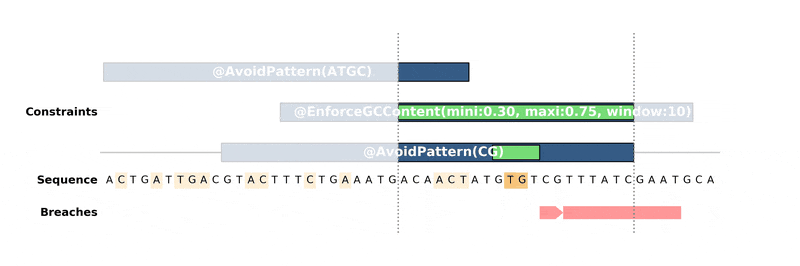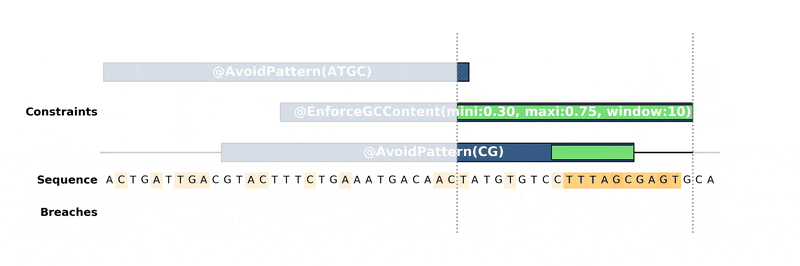
Installation¶
DNA Chisel requires Python 3, and can be installed via a pip command:
pip install dnachisel # <= minimal install without reports support
pip install 'dnachisel[reports]' # <= full install with all dependencies
The full installation using dnachisel[reports] downloads heavier libraries
(Matplotlib, PDF reports, sequenticon) for report generation, but is highly
recommended to use DNA Chisel interactively via Python scripts. Also install
GeneBlocks and its
dependencies if you wish to include a plot of sequence edits in the report.
Optionally, also install Bowtie to be able to use AvoidMatches (which
removes short homologies with existing genomes). On Ubuntu:
sudo apt-get install bowtie
License = MIT¶
DNA Chisel is an open-source software originally written at the Edinburgh Genome Foundry by Zulko and released on Github under the MIT licence (Copyright 2017 Edinburgh Genome Foundry, University of Edinburgh). Everyone is welcome to contribute!
More biology software¶

DNA Chisel is part of the EGF Codons synthetic biology software suite for DNA design, manufacturing and validation.
